LMU Newsroom
What is going on at LMU? Everything at a glance in the LMU Newsroom — news, events, interviews, backgrounds, stories.

ERC Advanced Grant for political scientist Christoph Knill
How does a permanent state of crisis affect policymaking? The European Research Council is funding an LMU project on this topic.
Read more
Cockayne syndrome: new insights into cellular DNA repair mechanism
LMU researchers decode repair mechanism during transcription of genetic information.
Read more
Hunting the bug
Computer scientist Marie-Christine Jakobs is an expert on the reliability of software.
Read moreINSIGHTS. Magazine

Polyglot machines
How artificial intelligence learns the rich variety of human languages: Hinrich Schütze, computational linguist at LMU, researches multilingual software that can do small languages. From the research magazine EINSICHTEN
Read more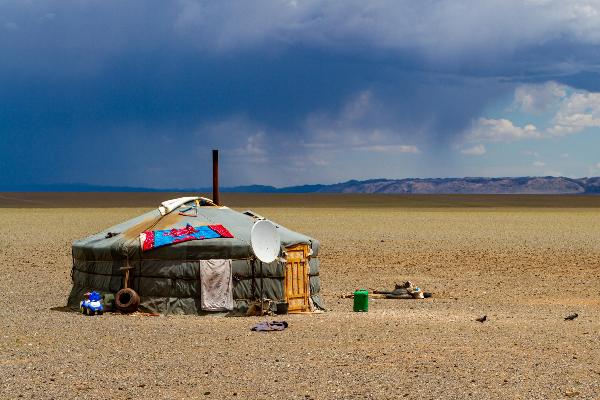
From the steppe to the city
A social transformation is underway in Mongolia, as many nomadic pastoralists move to urban areas. Geographer Lukas Lehnert investigates what this means for the environment and ecosystems. From the research magazine EINSICHTEN
Read more
“When success dictates self-worth”
When does wanting to do well become unwell? Barbara Cludius researches the propensity for perfectionism. In our EINSICHTEN interview she explains how a harmful thought construct is related to various mental disorders.
Read more