LMU Newsroom
What is going on at LMU? Everything at a glance in the LMU Newsroom — news, events, interviews, backgrounds, stories.

Studying - an experience that affects all areas of life
To get the lecture period of the new semester off to a good start, the LMU Community provides tips for successful studies.
Read more
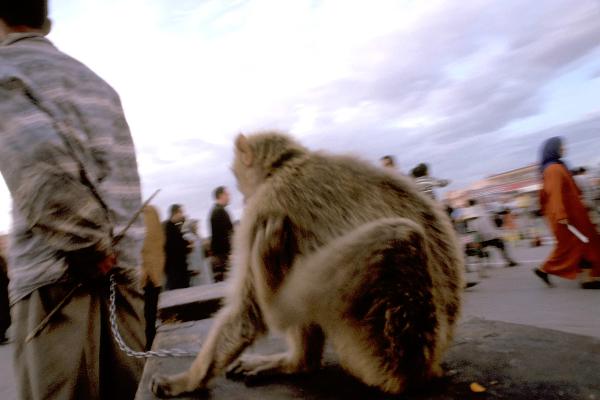
Creating the smallest building blocks for life
Physicist Erwin Frey researches the question, what is minimum for life.
Read more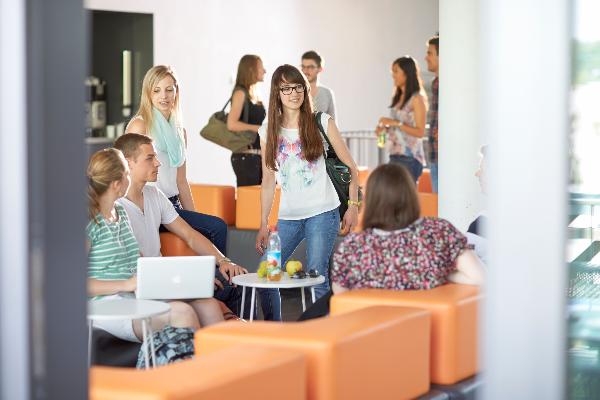
Making the world that much better
The contribution made by young people through volunteering is hugely important, especially in times of great social challenges.
Read moreINSIGHTS. Magazine

"Really?" - the new issue of INSIGHTS
"Echt jetzt" - the new issue of EINSICHTEN The new edition of the research magazine EINSICHTEN is out, with the main topic: "Really? Is the boundary between natural and artificial increasingly fading?" Click here for the highlights.
Read more
Polyglot machines
How artificial intelligence learns the rich variety of human languages: Hinrich Schütze, computational linguist at LMU, researches multilingual software that can do small languages. From the research magazine EINSICHTEN
Read more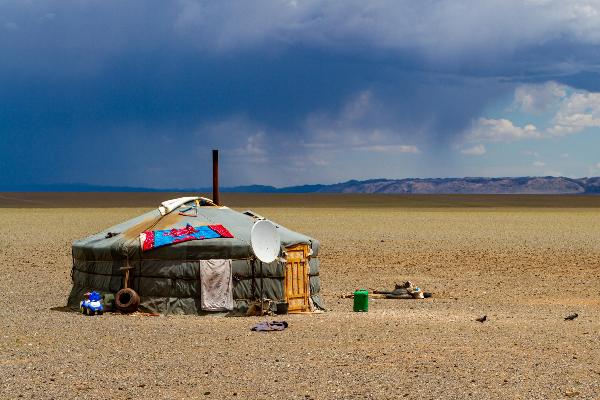
From the steppe to the city
A social transformation is underway in Mongolia, as many nomadic pastoralists move to urban areas. Geographer Lukas Lehnert investigates what this means for the environment and ecosystems. From the research magazine EINSICHTEN
Read more