LMU Newsroom
What is going on at LMU? Everything at a glance in the LMU Newsroom — news, events, interviews, backgrounds, stories.

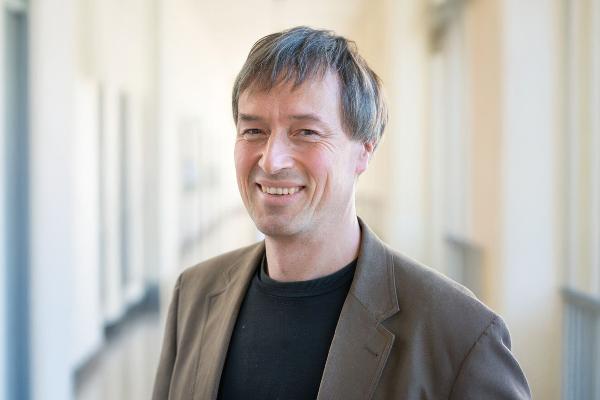
DFG funding for three LMU researchers
Neonatologist Vincent Gaertner, molecular biologist Marta Russo, and psychologist Alexander Soutschek receive funding from the Emmy Noether and Heisenberg programs.
Read more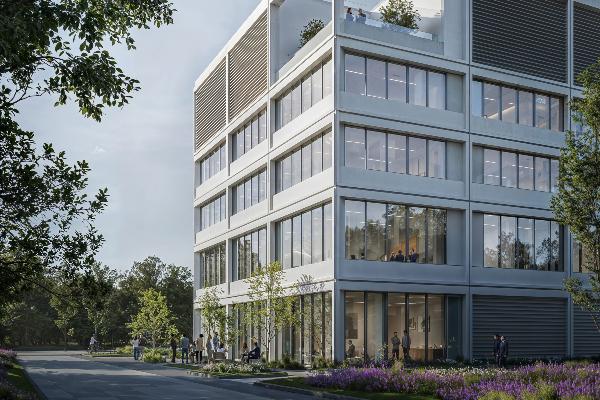
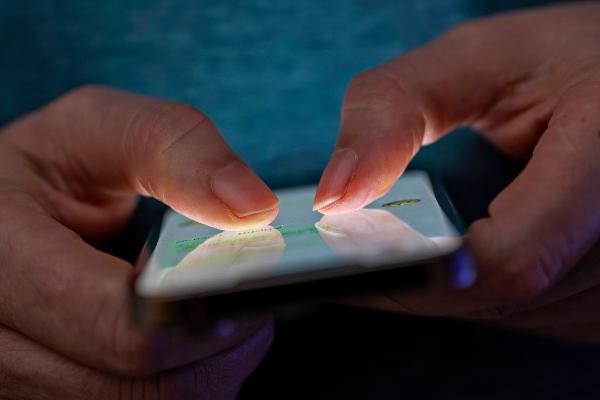
Research transfer: infection research leads to therapeutics
A cutting-edge center for immunology, infection, and pandemic research is being built in Penzberg. Scientific experts from LMU and LMU University Hospital will be playing a major role in the new research facility.
Read more
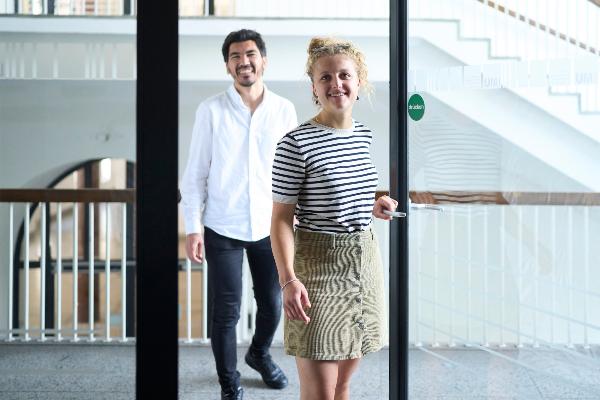
Foundations for innovations in higher education: new CRC at LMU
A new Collaborative Research Centre is being launched at LMU to investigate the possibilities of personalized and simulation-based learning at universities.
Read more